Featured Review: The Broad Footprint of Climate Change From Genes to Biomes to People ($)
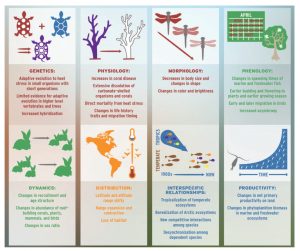 In this review, Scheffers et al. collate a vast amount of data to summarize the impact of climate change on terrestrial, freshwater and marine ecosystems; their findings are sobering. A key conclusion is that across the 94 processes analyzed, 84% show an impact of climate change. As one example, “plant phenologies were found to have shifted by more than two standard deviations across 54% of Earth’s land area during the past three decades.” The authors provide evidence of impacts on single species as well as interactions between species, for example through effects on competition, pollinator services and disease. The authors don’t pull any punches, and conclude, “Time is running out for a globally synchronized response to climate change that integrates adequate protection of biodiversity and ecosystem services”. Science 10.1126/science.aaf7671 Tags: Botany, Ecophysiology, Environmental Plant Biology
In this review, Scheffers et al. collate a vast amount of data to summarize the impact of climate change on terrestrial, freshwater and marine ecosystems; their findings are sobering. A key conclusion is that across the 94 processes analyzed, 84% show an impact of climate change. As one example, “plant phenologies were found to have shifted by more than two standard deviations across 54% of Earth’s land area during the past three decades.” The authors provide evidence of impacts on single species as well as interactions between species, for example through effects on competition, pollinator services and disease. The authors don’t pull any punches, and conclude, “Time is running out for a globally synchronized response to climate change that integrates adequate protection of biodiversity and ecosystem services”. Science 10.1126/science.aaf7671 Tags: Botany, Ecophysiology, Environmental Plant Biology
Featured Review: Evolution, Genomics and Epidemiology of Pseudomonas syringae ($)
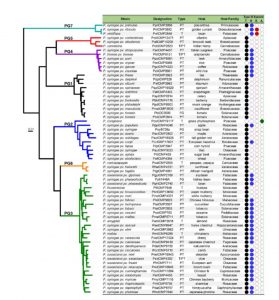 Much of our understanding of the molecular interactions between plants and bacterial pathogens comes from studies of the interaction between two model organisms: Arabidopsis thaliana and Pseudomonas syringae. Baltrus et al. explore the genetic, phenotypic and epidemiological diversity of P. syringae, including the challenges of resolving taxonomy in organisms that freely exchange genetic material. They argue that P. syringae has the potential to continue to provide insights into agriculturally relevant evolutionary questions given the species’ “diversity, ubiquity, agronomic significance and rich knowledge base”. Mol. Plant Pathol. 10.1111/mpp.12506 Tags: Biotic Interactions, Botany, Evolution, Genomics
Much of our understanding of the molecular interactions between plants and bacterial pathogens comes from studies of the interaction between two model organisms: Arabidopsis thaliana and Pseudomonas syringae. Baltrus et al. explore the genetic, phenotypic and epidemiological diversity of P. syringae, including the challenges of resolving taxonomy in organisms that freely exchange genetic material. They argue that P. syringae has the potential to continue to provide insights into agriculturally relevant evolutionary questions given the species’ “diversity, ubiquity, agronomic significance and rich knowledge base”. Mol. Plant Pathol. 10.1111/mpp.12506 Tags: Biotic Interactions, Botany, Evolution, Genomics
Featured Update: Virus-induced flowering, a breeding and research tool
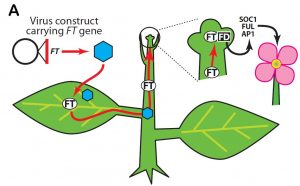 What’s good for the plant is not always good for the plant breeder. For example, the long juvenile phase that enables an orange tree to reach a good size before it invests energy into flower and fruit production means that it takes decades for a breeder’s work to come to fruition. Similarly, plants that require a certain temperature or daylength before they flower create difficulties. The identification of FT (aka florigen) as a flowering switch has paved the way for accelerated breeding efforts. McGarry et al. review the use of Virus-induced flowering, in which FT is transiently expressed from a viral cassette, as a tool for research and breeding. Plant Physiol. 10.1104/pp.16.01336 Tags: Applied Plant Biology, Botany, Development, Growth Regulation, Signals and Responses
What’s good for the plant is not always good for the plant breeder. For example, the long juvenile phase that enables an orange tree to reach a good size before it invests energy into flower and fruit production means that it takes decades for a breeder’s work to come to fruition. Similarly, plants that require a certain temperature or daylength before they flower create difficulties. The identification of FT (aka florigen) as a flowering switch has paved the way for accelerated breeding efforts. McGarry et al. review the use of Virus-induced flowering, in which FT is transiently expressed from a viral cassette, as a tool for research and breeding. Plant Physiol. 10.1104/pp.16.01336 Tags: Applied Plant Biology, Botany, Development, Growth Regulation, Signals and Responses
Improving photosynthesis by accelerating recovery from photoprotection
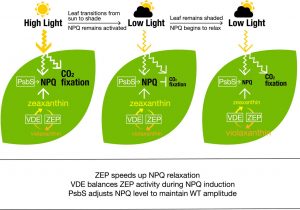 Plants have a mechanism to protect themselves from excess sunlight by harmlessly dissipating some of it as heat, by a process named nonphotochemical quenching of chlorophyll fluorescence (NPQ). But this process depresses the photosynthetic efficiency of these plants eventually affecting the overall crop yields. Kromdijk et al. describe the bioengineering of a faster response to natural shading events in Nicotiana (tobacco), resulting in an increased plant productivity by about 15% in fluctuating light. As this photoprotective mechanism is common to all crops, this study may provide insights to obtaining a sustainable increase in crop productivity and thereby higher crop yields in order to address the world’s future food requirements. (Aummary by Bhavisha P. Sheth) Science 10.1126/science.aai8878 Tags: Applied Plant Biology, Biochemistry, Bioenergetics, Ecophysiology, Physiology
Plants have a mechanism to protect themselves from excess sunlight by harmlessly dissipating some of it as heat, by a process named nonphotochemical quenching of chlorophyll fluorescence (NPQ). But this process depresses the photosynthetic efficiency of these plants eventually affecting the overall crop yields. Kromdijk et al. describe the bioengineering of a faster response to natural shading events in Nicotiana (tobacco), resulting in an increased plant productivity by about 15% in fluctuating light. As this photoprotective mechanism is common to all crops, this study may provide insights to obtaining a sustainable increase in crop productivity and thereby higher crop yields in order to address the world’s future food requirements. (Aummary by Bhavisha P. Sheth) Science 10.1126/science.aai8878 Tags: Applied Plant Biology, Biochemistry, Bioenergetics, Ecophysiology, Physiology
Nramp5 mediates uptake of Cadmium and Manganese in Barley
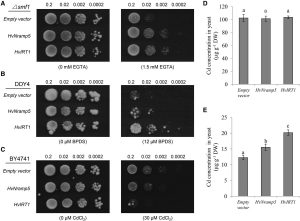 The Natural Resistance Associated Macrophage Protein (Nramp) family of transporters is mainly responsible for uptake of metal ions in all organisms including bacteria, animals, and plants. Some members of this family in plants have been functionally characterized, especially in model plants such as Arabidopsis and rice but no members of Nramp family has been characterized in barley (Hordeum vulgare) so far. Wu et al. isolated and functionally characterized HvNramp5 and show that it transports Mn2+ and Cd2+, but not Fe2+ in yeast. They have also shown that HvNramp5 is mainly localized at the epidermal cells of the root tips and knockdown of HvNramp5 resulted in decreased accumulation of Mn and Cd, but not Fe, Cu, and Zn in both roots and shoots. (Summary by Mather Khan) Plant Physiol. 10.1104/pp.16.01189 Tags: Abiotic Interactions, Biochemistry, Physiology, Transport
The Natural Resistance Associated Macrophage Protein (Nramp) family of transporters is mainly responsible for uptake of metal ions in all organisms including bacteria, animals, and plants. Some members of this family in plants have been functionally characterized, especially in model plants such as Arabidopsis and rice but no members of Nramp family has been characterized in barley (Hordeum vulgare) so far. Wu et al. isolated and functionally characterized HvNramp5 and show that it transports Mn2+ and Cd2+, but not Fe2+ in yeast. They have also shown that HvNramp5 is mainly localized at the epidermal cells of the root tips and knockdown of HvNramp5 resulted in decreased accumulation of Mn and Cd, but not Fe, Cu, and Zn in both roots and shoots. (Summary by Mather Khan) Plant Physiol. 10.1104/pp.16.01189 Tags: Abiotic Interactions, Biochemistry, Physiology, Transport
Control of seed dormancy by antisense ncRNA ($)
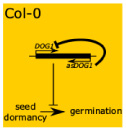 Seed dormancy, a block in the ability of seeds to germinate under otherwise favorable conditions, is tightly controlled by the Delay of Germination 1 gene (DOG1). As with many other genes that regulate important life cycle transitions, DOG1 expression is highly regulated in many ways. In this paper, Fedak et al. show yet another layer of regulation of DOG1 expression that involves the production of an antisense noncoding transcript (asDOG1). The authors observed that low asDOG1 levels correlate with increased amounts of sense DOG1 mRNA, and, in turn, with a deep dormant state in seeds. As the promoter for this antisense transcript is highly conserved in the Brassicaceae family, this mechanism would be expected to be active in other plants. (Summary by Gaby Auge) Proc. Natl. Acad. Sci. USA, doi:10.1073/pnas.1608827113 Tags: Development, Gene Regulation, Genetics, Molecular Biology
Seed dormancy, a block in the ability of seeds to germinate under otherwise favorable conditions, is tightly controlled by the Delay of Germination 1 gene (DOG1). As with many other genes that regulate important life cycle transitions, DOG1 expression is highly regulated in many ways. In this paper, Fedak et al. show yet another layer of regulation of DOG1 expression that involves the production of an antisense noncoding transcript (asDOG1). The authors observed that low asDOG1 levels correlate with increased amounts of sense DOG1 mRNA, and, in turn, with a deep dormant state in seeds. As the promoter for this antisense transcript is highly conserved in the Brassicaceae family, this mechanism would be expected to be active in other plants. (Summary by Gaby Auge) Proc. Natl. Acad. Sci. USA, doi:10.1073/pnas.1608827113 Tags: Development, Gene Regulation, Genetics, Molecular Biology
ABI5 is a Regulator of Seed Maturation and Longevity in Legumes ($)
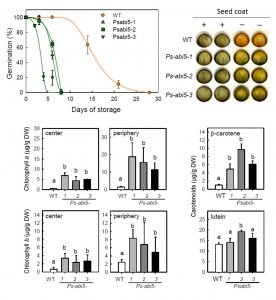 The ability of seeds to remain viable in storage is crucial for conservation of germplasm resources as well as to ensure germination of newly-planted seeds. Through a quantitative-trait locus (QTL) study in Medicago truncatula, Zinsmeister et al. identified the transcription factor ABI5 as a determinant of seed longevity. The abi5 mutant seeds showed altered patterns of expression of genes associated with maturation including LEA genes, and were impaired in chlorophyll breakdown. The significance of chlorophyll retention and its contributiton to decreased seed longevity is discussed. Plant Cell 0.1105/tpc.16.00470 Tags: Applied Plant Biology, Biochemistry, Gene Regulation, Genetics, Metabolism, Physiology
The ability of seeds to remain viable in storage is crucial for conservation of germplasm resources as well as to ensure germination of newly-planted seeds. Through a quantitative-trait locus (QTL) study in Medicago truncatula, Zinsmeister et al. identified the transcription factor ABI5 as a determinant of seed longevity. The abi5 mutant seeds showed altered patterns of expression of genes associated with maturation including LEA genes, and were impaired in chlorophyll breakdown. The significance of chlorophyll retention and its contributiton to decreased seed longevity is discussed. Plant Cell 0.1105/tpc.16.00470 Tags: Applied Plant Biology, Biochemistry, Gene Regulation, Genetics, Metabolism, Physiology
Conservation of tip-growth genes from liverwort to Arabidopsis
 Marchantia polymorpha, a liverwort, diverged more than 460 million years ago from the vascular plant lineage. Comparisons between vascular plants and liverworts shed light onto their common ancestor, which was an early land plant. Honkanan et al. generated and screened 336,000 M. polymorpha T-DNA insertion mutants from which they identified 33 genes required for normal rhizoid formation. They also generated a draft M. polymorpha genome assembly onto which they mapped the T-DNA insertions. Many of the genes they identified as necessary for rhizoid formation and tip growth are also expressed in or functional in Arabidopsis root hairs, showing that the mechanism of tip-growth has been largely conserved for more than 460 million years. Curr. Biol. 10.1016/j.cub.2016.09.062 Tags: Botany, Cell Biology, Development, Evolution, Genetics
Marchantia polymorpha, a liverwort, diverged more than 460 million years ago from the vascular plant lineage. Comparisons between vascular plants and liverworts shed light onto their common ancestor, which was an early land plant. Honkanan et al. generated and screened 336,000 M. polymorpha T-DNA insertion mutants from which they identified 33 genes required for normal rhizoid formation. They also generated a draft M. polymorpha genome assembly onto which they mapped the T-DNA insertions. Many of the genes they identified as necessary for rhizoid formation and tip growth are also expressed in or functional in Arabidopsis root hairs, showing that the mechanism of tip-growth has been largely conserved for more than 460 million years. Curr. Biol. 10.1016/j.cub.2016.09.062 Tags: Botany, Cell Biology, Development, Evolution, Genetics
Evolution of small-petal size in a recently diverged self-fertilizing species ($)
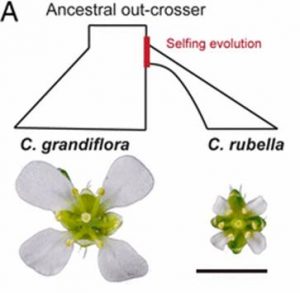 Capsella grandiflora is an out-crossing species from which diverged a self-fertilizing species Capsella rubella within the last 200,000 years (recent in evolutionary terms). Large petals like those in C. grandiflora attract pollinators, whereas there is some evidence that smaller petals like those in C. rubella better support self-fertilization. Sicard et al. find genetic evidence for the rapid and specific decrease in petal size in C. rubella, in the form of a tissue-specific enhancer in the intron of a growth-associated gene. Further, they show that C. rubella’s small-petal allele arose from capture and fixation of standing-genetic variation within the ancestral population. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1613394113 Tags: Botany, Development, Evolution, Genetics, Growth Regulation
Capsella grandiflora is an out-crossing species from which diverged a self-fertilizing species Capsella rubella within the last 200,000 years (recent in evolutionary terms). Large petals like those in C. grandiflora attract pollinators, whereas there is some evidence that smaller petals like those in C. rubella better support self-fertilization. Sicard et al. find genetic evidence for the rapid and specific decrease in petal size in C. rubella, in the form of a tissue-specific enhancer in the intron of a growth-associated gene. Further, they show that C. rubella’s small-petal allele arose from capture and fixation of standing-genetic variation within the ancestral population. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1613394113 Tags: Botany, Development, Evolution, Genetics, Growth Regulation
Insights into domestication of maize from 5,310-year-old ancient DNA
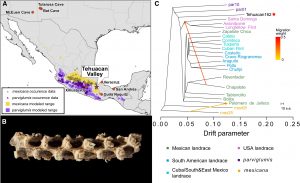 Ramos-Madrigal et al. sequenced the genome of a 5,310-year-old maize cob, known as Tehuacan162, excavated in the 1960s from the Tehuacan Valley of Mexico. They compared the Tehuacan162 genome to the genomes of modern landraces, and to the closest living relatives of maize, Zea maize ssp. parviglumis and Zea maize ssp. mexicana. Their data show that Tehuacan162 is an ancient form of maize closely related to the ancestor of all modern maize varieties and represents an early stage of domestication. Some but not all domestication-associated genes remain in the ancestral state, suggesting that much of maize’s evolution during domestication may have been more gradual than previously thought. (Summary by K.K. Sabu) Curr. Biol. 10.1016/j.cub.2016.09.036 Tags: Botany, Evolution
Ramos-Madrigal et al. sequenced the genome of a 5,310-year-old maize cob, known as Tehuacan162, excavated in the 1960s from the Tehuacan Valley of Mexico. They compared the Tehuacan162 genome to the genomes of modern landraces, and to the closest living relatives of maize, Zea maize ssp. parviglumis and Zea maize ssp. mexicana. Their data show that Tehuacan162 is an ancient form of maize closely related to the ancestor of all modern maize varieties and represents an early stage of domestication. Some but not all domestication-associated genes remain in the ancestral state, suggesting that much of maize’s evolution during domestication may have been more gradual than previously thought. (Summary by K.K. Sabu) Curr. Biol. 10.1016/j.cub.2016.09.036 Tags: Botany, Evolution
Origins of potato domestication ($)
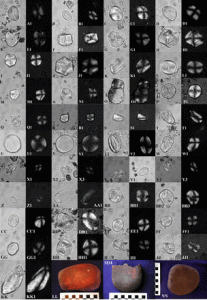 Evidence in an early village in the Titicaca Basin in Perú, Jiskairumoko, shows the transition to sedentism and food production, and points to south-central Andes as the origin of potato domestication, among many other crops. Rumold and Aldenderfer show starch microbotanical evidence that directly correlates with exploitation, and probably early cultivation, of potato in said site. The microremains analyzed suggest that by ~3400-1600 calendar years before present, the residents either had already selected (and domesticated) potato plants that had reduced toxicity by glycoalkaloids, or that they found a way to detox and make use of wild Solanum (by grinding, cooking or adding comestible earths). Although they cannot ascertain whether the analyzed Solanum starches from the site derive from cultivated/domesticated or wild potato, the data presented here shed light in the origins of this staple food. (Paper summary by Gaby Auge) Proc. Natl. Acad. Sci. USA doi:10.1073/pnas.1604265113 Tags: Applied Plant Biology, Botany
Evidence in an early village in the Titicaca Basin in Perú, Jiskairumoko, shows the transition to sedentism and food production, and points to south-central Andes as the origin of potato domestication, among many other crops. Rumold and Aldenderfer show starch microbotanical evidence that directly correlates with exploitation, and probably early cultivation, of potato in said site. The microremains analyzed suggest that by ~3400-1600 calendar years before present, the residents either had already selected (and domesticated) potato plants that had reduced toxicity by glycoalkaloids, or that they found a way to detox and make use of wild Solanum (by grinding, cooking or adding comestible earths). Although they cannot ascertain whether the analyzed Solanum starches from the site derive from cultivated/domesticated or wild potato, the data presented here shed light in the origins of this staple food. (Paper summary by Gaby Auge) Proc. Natl. Acad. Sci. USA doi:10.1073/pnas.1604265113 Tags: Applied Plant Biology, Botany
Climate-driven leaf-out shifts in woody plants ($)
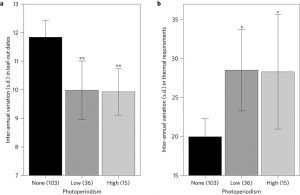 Climate change is causing earlier leaf emergence times in trees (dormancy release) as springs become warmer. However, there are other cues, like photoperiod and chilling, that may hinder this linear response to warming and influence species ranges. From a study with 173 tree species from a broad native range in the Northern Hemisphere, Zohner et al. observed that reduced chilling times increased sensitivity requirements to photoperiod. This was especially true for species from lower latitudes (shorter winters), while species from regions with long winters are likely to rely on signals such us winter chilling and spring temperature, more than day length. These results suggest that photoperiod may constrain northward range expansion for many species even if facing warming environments. (Summary by Gaby Auge) Nature Climate Change, doi:10.1038/nclimate3138 Tags: Abiotic Interactions, Botany, Ecophysiology, Environmental Plant Biology
Climate change is causing earlier leaf emergence times in trees (dormancy release) as springs become warmer. However, there are other cues, like photoperiod and chilling, that may hinder this linear response to warming and influence species ranges. From a study with 173 tree species from a broad native range in the Northern Hemisphere, Zohner et al. observed that reduced chilling times increased sensitivity requirements to photoperiod. This was especially true for species from lower latitudes (shorter winters), while species from regions with long winters are likely to rely on signals such us winter chilling and spring temperature, more than day length. These results suggest that photoperiod may constrain northward range expansion for many species even if facing warming environments. (Summary by Gaby Auge) Nature Climate Change, doi:10.1038/nclimate3138 Tags: Abiotic Interactions, Botany, Ecophysiology, Environmental Plant Biology

Vuestros artículos son muy interesantes.
A great selection of articles, which I enjoyed reading. This is a really valuable aid for busy people; like me!
Thanks, it’s great to hear from you. Do let us know if you see a paper we should include at any time!