This week’s Research in Focus is reprinted from an In Brief published by Science Editor Nancy Hofmann in The Plant Cell, which summarizes an article newly published by Calderwood et al. This study explores the distribution and identity of mobile mRNA within the plant body, and presents a hypothesis that abundance in companion cells is a key determinant of mRNA mobility.
Messenger RNA (mRNA) can be transported long distances within a plant via the phloem sap. Specific mobile transcripts are reported to influence flowering as well as leaf, tuber, and root development, and mobile mRNAs have been suggested to serve as long-distance signaling molecules within the plant (reviewed in Spiegelman et al., 2013). Among the several questions surrounding long-distance transport of mRNAs is what mechanism determines which transcripts are mobile. A recent report indicates that a much higher proportion of transcripts are mobile than previously thought. Thieme et al. (2015) grafted two different ecotypes of Arabidopsis thaliana and identified more than 2000 genes corresponding to mobile mRNAs. Such widespread trafficking poses further questions about what role these mobile transcripts might play in the plant, as well as the degree to which their mobility is sequence specific.
A new Breakthrough Report from Calderwood et al. (2016) takes advantage of the availability of this large data set of mobile transcripts to test whether mRNA mobility can be explained by the abundance of that transcript in companion cells. The authors built a computational model in which individual mRNA molecules carry out a random walk through a box-shaped companion cell (See Figure). At each step, the mRNA moves a set distance in any direction or decays, the latter possibility occurring with a set probability. Upon reaching the cell surface, the mRNA has a set (transcript-independent) probability of entering the sieve element for transport.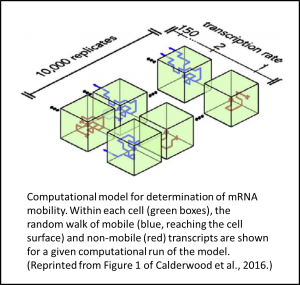
The effect of mRNA abundance on mRNA mobility (i.e., the likelihood of the transcript to leave the companion cell) was tested in the model by varying the initial concentration of a given mRNA (to model changes in transcription rate) and its probability of decay (to model differences in half-life). In this model, the probability of mobility would eventually saturate with increasing mRNA abundance. To test the model, the authors compared its prediction regarding mobility relative to abundance to the experimental data provided by Thieme et al. (2015). The distribution of experimentally determined mobile and non-mobile transcripts with abundance was largely consistent with the prediction of the model.
As decay is an important factor in determination of mRNA abundance, the authors explored whether stable transcripts were more likely to be mobile. Indeed, the half-life of the mobile transcripts was longer than that of nonmobile transcripts. In addition, longer transcripts were less likely to be mobile, consistent with the idea that larger molecules would have a slower diffusion rate through the cell and thus a lower likelihood of reaching the plasmodesmata at the cell surface.
Although the success of the model suggests that abundance in the companion cell is the major determinant of mobility, the model did not correctly predict mobility for some mRNAs at the low end of the abundance distribution. These low-abundance transcripts were enriched for three sequence motifs, which could point to a sequence-specific mechanism of mobility for this set of transcripts. The diffusion model does not explain the tissue-specific allocation of hundreds of mRNAs observed to move in Arabidopsis. However, the ability of the abundance model to explain the mobility of a large population of mRNA suggests that most detected mRNA mobility is non-sequence specific, at least in terms of leaving the companion cell to enter the phloem sap. This does not preclude the possibility of active regulation of mRNA mobility at the level of abundance in the companion cell, and it does not rule out sequence-specific mechanisms for trafficking of certain mRNAs. It does indicate, however, that an abundance-driven determination of mobility should probably be the null hypothesis for any mRNA found to be trafficked in the phloem.
REFERENCES
Calderwood, A., Kopriva, S., and Morris, R.J. (2016). Transcript abundance explains mRNA mobility data in Arabidopsis thaliana. Plant Cell. doi: 10.1105/tpc.15.00956.
Spiegelman, Z., Golan, G., and Wolf, S. (2013). Don’t kill the messenger: Long-distance trafficking of mRNA molecules. Plant Sci. 213: 1-8.
Thieme, C.J., Rojas-Triana, M., Stecyk, E., Schudoma, C., Zhang, W., Yang, L., Miñambres, M., Walther, D., Schulze, W.X., Paz-Ares, J., Scheible, W.-R., and Kragler, F. (2015). Endogenous Arabidopsis messenger RNAs transported to distant tissues. Nature Plants 1: 15025.
