Featured Review: Role of Mass Spectrometry in Biology
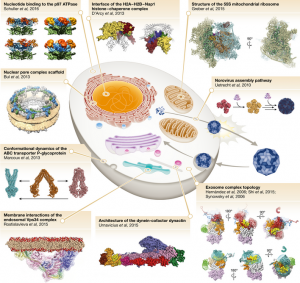 Mass spectrometry (MS) has become a core tool for molecular and structural biology. Lössl et al. review the history of MS and provide an accessible explanation for how it works and its applications, with numerous examples. This is a must-read for students of biochemistry as well as practicing scientists. EMBO J. 10.15252/embj.201694818 Tags: Biochemistry, Computational Biology, Molecular Biology
Mass spectrometry (MS) has become a core tool for molecular and structural biology. Lössl et al. review the history of MS and provide an accessible explanation for how it works and its applications, with numerous examples. This is a must-read for students of biochemistry as well as practicing scientists. EMBO J. 10.15252/embj.201694818 Tags: Biochemistry, Computational Biology, Molecular Biology
Featured Review: Improving yield potential in cassava
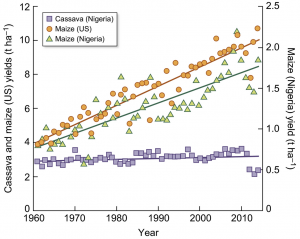 Cassava (Manihot esculenta) is a vital food source for millions living on the African continent, but cassava yield increases have not kept pace with other staple crops. DeSouza et al. investigate the efficiency of light interception and light conversion into biomass in cassava; both values fall well below theoretical values, indicating possibilities for yield improvement. The review concludes with a list of opportunities for trait improvement (e.g., alterations in canopy architecture and leaf angle) as well as a their expected efficiency gains and timescales. New Phytol. 10.1111/nph.14250 Tags: Applied Plant Biology, Bioenergetics, Ecophysiology, Physiology
Cassava (Manihot esculenta) is a vital food source for millions living on the African continent, but cassava yield increases have not kept pace with other staple crops. DeSouza et al. investigate the efficiency of light interception and light conversion into biomass in cassava; both values fall well below theoretical values, indicating possibilities for yield improvement. The review concludes with a list of opportunities for trait improvement (e.g., alterations in canopy architecture and leaf angle) as well as a their expected efficiency gains and timescales. New Phytol. 10.1111/nph.14250 Tags: Applied Plant Biology, Bioenergetics, Ecophysiology, Physiology
 Featured Review: How many ways are there to make a root? ($)
Featured Review: How many ways are there to make a root? ($)
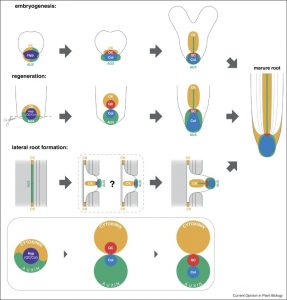
Different roots have different origins: the primary root forms very early in the development of the embryo, whereas lateral roots come from pericycle cells, adventitious roots can come from a variety of vasculature-associated cell types, and damaged-induced roots have yet other origins. At some point, no matter where the roots begin, their developmental progressions converge. Birnbaum reviews the common and divergent mechanisms of the earliest stages of root formation. Curr. Opin. Plant Biol. 10.1016/j.pbi.2016.10.001 Tags: Development, Growth Regulation, Signals and Responses
See also:
Establishment of transcriptional cascades in roots ($)
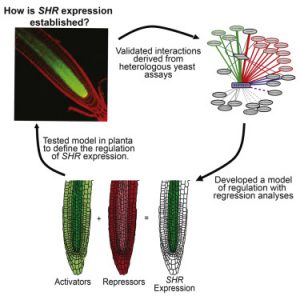 The establishment of cellular differentiation patterns in multicellular organisms relies on suites of transcription factors that specify cell fates. SHORTROOT (SHR) and SCARECROW (SCR) are two interdependent transcription factors that set up root ground tissues. Forward genetic approaches have failed to identify factors upstream of SHR. Sparks et al. take a systems approach (involving analysis of cell-type specific transcriptomic data and enhanced yeast-one-hybrid assays) to identify networks of interacting activators and repressors that establish the SHR-SCR transcriptional cascade. Devel. Cell. 10.1016/j.devcel.2016.09.031 Tags: Computational Biology, Development, Gene Regulation, Molecular Biology, Systems Biology
The establishment of cellular differentiation patterns in multicellular organisms relies on suites of transcription factors that specify cell fates. SHORTROOT (SHR) and SCARECROW (SCR) are two interdependent transcription factors that set up root ground tissues. Forward genetic approaches have failed to identify factors upstream of SHR. Sparks et al. take a systems approach (involving analysis of cell-type specific transcriptomic data and enhanced yeast-one-hybrid assays) to identify networks of interacting activators and repressors that establish the SHR-SCR transcriptional cascade. Devel. Cell. 10.1016/j.devcel.2016.09.031 Tags: Computational Biology, Development, Gene Regulation, Molecular Biology, Systems Biology
Combinatorial interaction network of nitrogen and hormones ($)
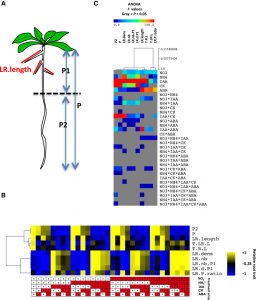 Roots respond to exogenous and endogenous factors by altering patterns of gene expression and growth. Ristova at al. examine the combinatorial effects of nitrogen-containing nutrients (nitrate and ammonium) and hormones (auxin, cytokinin, and abscisic acid). From these data they develop a model that reveals both early and late gene responses and identifies genes that may play key roles in the integration of these signals. Science Signaling 10.1126/scisignal.aaf2768 Tags: Computational Biology, Development, Gene Regulation, Systems Biology
Roots respond to exogenous and endogenous factors by altering patterns of gene expression and growth. Ristova at al. examine the combinatorial effects of nitrogen-containing nutrients (nitrate and ammonium) and hormones (auxin, cytokinin, and abscisic acid). From these data they develop a model that reveals both early and late gene responses and identifies genes that may play key roles in the integration of these signals. Science Signaling 10.1126/scisignal.aaf2768 Tags: Computational Biology, Development, Gene Regulation, Systems Biology
Featured Review: Molecular Genetics of Grass Stomata ($)
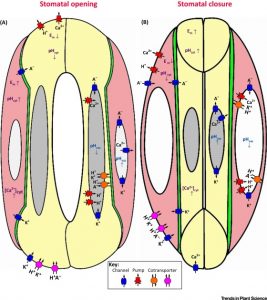 The ability to control the movement of water and gases through stomatal pores was crucial in the adaptation of plants to dry land. The precision with which the stomatal complex controls transpiration varies amongst plants, with grasses showing the greatest level of control. Chen et al. review the diversity of stomatal structure and function across plant species, to recapitulate the events leading to the evolution of grass stomata. Trends Plant Sci. 10.1016/j.tplants.2016.09.005 Tags: Botany, Cell Biology, Development, Ecophysiology, Transport
The ability to control the movement of water and gases through stomatal pores was crucial in the adaptation of plants to dry land. The precision with which the stomatal complex controls transpiration varies amongst plants, with grasses showing the greatest level of control. Chen et al. review the diversity of stomatal structure and function across plant species, to recapitulate the events leading to the evolution of grass stomata. Trends Plant Sci. 10.1016/j.tplants.2016.09.005 Tags: Botany, Cell Biology, Development, Ecophysiology, Transport
See also:
ABA controlled sex before transpiration in plant evolution($)
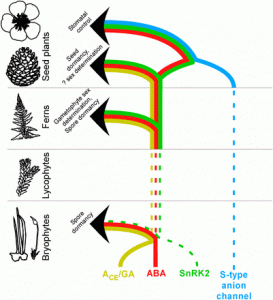 McAdam et al. show that the hormone abscisic acid (ABA) contributes to fern sexual differentiation by regulating the ratio of male to hermaphrodite tissues. They trace the evolutionary history of the ABA-SNRK2 signaling pathway from determining the sex of ferns to controlling seed dormancy and transpiration. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1606614113 Tags: Botany, Development, Evolution, Signals and Responses
McAdam et al. show that the hormone abscisic acid (ABA) contributes to fern sexual differentiation by regulating the ratio of male to hermaphrodite tissues. They trace the evolutionary history of the ABA-SNRK2 signaling pathway from determining the sex of ferns to controlling seed dormancy and transpiration. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1606614113 Tags: Botany, Development, Evolution, Signals and Responses
Featured Review: Horizontal gene transfer in parasitic plants ($)
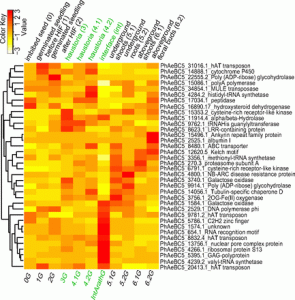 Horizontal gene transfer (HGT), or the non-sexual movement of genes from one organism to another, is a widespread phenomenon in prokaryotes; for example, HGT is responsible for the spread of antibiotic resistance through a bacterial population. HGT is less common between eukaryotes, but HGT is known to occur between host and parasitic plant. Yang et al. identified more than 50 HGT events from host to parasite. Their analysis shows that transferred genes are preferentially expressed in the haustrorium (the feeding structure of parasitic plants) and that they show evidence of positive selection, suggesting that these genes are benefitting the parasitic plant. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1608765113 Tags: Biotic interactions, Botany, Evolution
Horizontal gene transfer (HGT), or the non-sexual movement of genes from one organism to another, is a widespread phenomenon in prokaryotes; for example, HGT is responsible for the spread of antibiotic resistance through a bacterial population. HGT is less common between eukaryotes, but HGT is known to occur between host and parasitic plant. Yang et al. identified more than 50 HGT events from host to parasite. Their analysis shows that transferred genes are preferentially expressed in the haustrorium (the feeding structure of parasitic plants) and that they show evidence of positive selection, suggesting that these genes are benefitting the parasitic plant. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1608765113 Tags: Biotic interactions, Botany, Evolution
Featured Review: Biology and evolution of arbuscular mycorrhizal symbiosis in the light of genomics
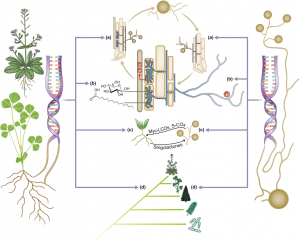 Most plants associate with arbuscular mycorrhizal fungi in a symbiosis that benefits both partners. The origins and benefits of the interactions between arbuscular mycrorrhizal fungi and plants is becoming clearer as additional genome sequences are unraveled. Kamel et al. review our understanding of this extraordinary partnership and the insights provided by the genomic sequences of both host and nonhost plants as well as their fungal partners. New Phytol. Tags: Biotic interactions, Botany, Evolution, Genomics
Most plants associate with arbuscular mycorrhizal fungi in a symbiosis that benefits both partners. The origins and benefits of the interactions between arbuscular mycrorrhizal fungi and plants is becoming clearer as additional genome sequences are unraveled. Kamel et al. review our understanding of this extraordinary partnership and the insights provided by the genomic sequences of both host and nonhost plants as well as their fungal partners. New Phytol. Tags: Biotic interactions, Botany, Evolution, Genomics
Integration of light and temperature cues by phytochrome B ($)
 Although temperature is one of the most important cues that determine plant growth and development, with the predicted consequences that changes in temperature (climate change) might have on plant phenology and distribution, the identification of a thermosensor in plants has been elusive… but only until last week. Two groups have independently demonstrated that the red/far-red light photoreceptor phytochrome B (phyB) functions as a thermosensor in Arabidopsis. Legris et al. have shown that the light-independent inactivation of phyB is dependent on temperature, either if seedlings were exposed to light or kept in darkness, affecting the formation of nuclear bodies and probably modifying the expression of light- and temperature-regulated genes. On the other hand, Jung et al. have confirmed that phyB (and probably other phytochromes as well) controls expression of key temperature-responsive genes and, for some of them, it does so by directly binding to their promoters. Altogether, these results show that light and temperature signals are integrated by phyB, allowing plants to optimize responses to complex environments. Science: Legris et al. 10.1126/science.aaf5656 Jung et al. 10.1126/science.aaf6005 (Summary by Gabriela Auge) Tags: Abiotic Interactions, Cell Biology, Gene Regulation, Molecular Biology, Signals and Responses
Although temperature is one of the most important cues that determine plant growth and development, with the predicted consequences that changes in temperature (climate change) might have on plant phenology and distribution, the identification of a thermosensor in plants has been elusive… but only until last week. Two groups have independently demonstrated that the red/far-red light photoreceptor phytochrome B (phyB) functions as a thermosensor in Arabidopsis. Legris et al. have shown that the light-independent inactivation of phyB is dependent on temperature, either if seedlings were exposed to light or kept in darkness, affecting the formation of nuclear bodies and probably modifying the expression of light- and temperature-regulated genes. On the other hand, Jung et al. have confirmed that phyB (and probably other phytochromes as well) controls expression of key temperature-responsive genes and, for some of them, it does so by directly binding to their promoters. Altogether, these results show that light and temperature signals are integrated by phyB, allowing plants to optimize responses to complex environments. Science: Legris et al. 10.1126/science.aaf5656 Jung et al. 10.1126/science.aaf6005 (Summary by Gabriela Auge) Tags: Abiotic Interactions, Cell Biology, Gene Regulation, Molecular Biology, Signals and Responses
A robust gene-stacking method utilizing yeast assembly for plant synthetic biology
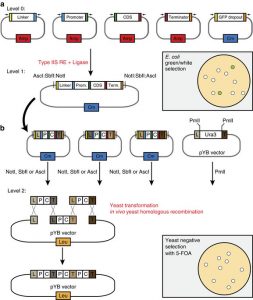 To efficiently engineer and reprogram plants to express new combinations of traits, such as disease resistance or biochemical pathways, versatile and efficient tools must be developed for synthetic biology. Shih et al. describe the development of a gene stacking method that takes advantage of homologous recombination in yeast to assemble large DNA vectors for plant transformation. Nature Comm. 10.1038/ncomms13215 Tags: Synthetic Biology (Congratulations to all the planty iGEM teams who presented at the iGEM jamboree, go team green!)
To efficiently engineer and reprogram plants to express new combinations of traits, such as disease resistance or biochemical pathways, versatile and efficient tools must be developed for synthetic biology. Shih et al. describe the development of a gene stacking method that takes advantage of homologous recombination in yeast to assemble large DNA vectors for plant transformation. Nature Comm. 10.1038/ncomms13215 Tags: Synthetic Biology (Congratulations to all the planty iGEM teams who presented at the iGEM jamboree, go team green!)
Rapid gene isolation in barley and wheat by mutant chromosome sequencing
 The large genome sizes of wheat and barley make it difficult to map and clone phenotypically-identified genes. Sánchez-Martín et al. present a new method, MutChromSeq, that makes gene mapping much more feasible. Their method starts with a conventional approach and the mapping of the locus-of-interest to a specific chromosome. Next, chromosomes are separated by chromosome flow sorting, to purify only the chromosome of interest. This greatly reduces the amount of DNA that must then be sequenced, greatly facilitating the process of gene identification. Genome Biol. 10.1186/s13059-016-1082-1 Tags: Applied Plant Biology, Genetics
The large genome sizes of wheat and barley make it difficult to map and clone phenotypically-identified genes. Sánchez-Martín et al. present a new method, MutChromSeq, that makes gene mapping much more feasible. Their method starts with a conventional approach and the mapping of the locus-of-interest to a specific chromosome. Next, chromosomes are separated by chromosome flow sorting, to purify only the chromosome of interest. This greatly reduces the amount of DNA that must then be sequenced, greatly facilitating the process of gene identification. Genome Biol. 10.1186/s13059-016-1082-1 Tags: Applied Plant Biology, Genetics
Genome of maize elite inbred line PH207
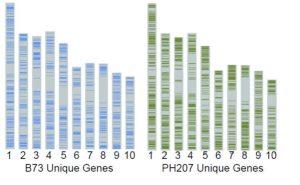 Zea mays has been one of the first and most important experimental organisms for geneticists and also is one of our most important food sources. Until now, B73 was the only publicly available complete maize whole genome assembly. Hirsch et al. have now released a draft genome assembly of a second inbred line, PH207, that is widely used in the production of hybrid seed. They also present a comparison of the transcript profiles of the two lines. The comparisons between these two datasets are particularly interesting given that both genotypes have been under intensive breeding selection for more than 100 years. Plant Cell 10.1105/tpc.16.00353 Tags: Applied plant biology, Computational Biology, Evolution, Genetics, Genomics
Zea mays has been one of the first and most important experimental organisms for geneticists and also is one of our most important food sources. Until now, B73 was the only publicly available complete maize whole genome assembly. Hirsch et al. have now released a draft genome assembly of a second inbred line, PH207, that is widely used in the production of hybrid seed. They also present a comparison of the transcript profiles of the two lines. The comparisons between these two datasets are particularly interesting given that both genotypes have been under intensive breeding selection for more than 100 years. Plant Cell 10.1105/tpc.16.00353 Tags: Applied plant biology, Computational Biology, Evolution, Genetics, Genomics
Genome of Cardamine hirsuta ($)
 Cardamine hirsuta, a small plant closely related to Arabidopsis thaliana, has been developed as a genetic model and has found particular success in the study of leaf development. Gan et al. release a high-quality reference genome for C. hirsuta that they compare to that of A. thaliana and other close relatives including A. lyrata and Capsella rubella. As a demonstration of the power of comparative genomics, they identify a tandem gene duplication in the C. hirsuta genome that distinguishes its explosive seed pod from those of its relatives. Nature Plants 10:1038/nplants.2016.167 Tags: Computational Biology, Development, Evolution, Genetics, Genomics
Cardamine hirsuta, a small plant closely related to Arabidopsis thaliana, has been developed as a genetic model and has found particular success in the study of leaf development. Gan et al. release a high-quality reference genome for C. hirsuta that they compare to that of A. thaliana and other close relatives including A. lyrata and Capsella rubella. As a demonstration of the power of comparative genomics, they identify a tandem gene duplication in the C. hirsuta genome that distinguishes its explosive seed pod from those of its relatives. Nature Plants 10:1038/nplants.2016.167 Tags: Computational Biology, Development, Evolution, Genetics, Genomics
Plants nanobionics for environmental sensing ($)
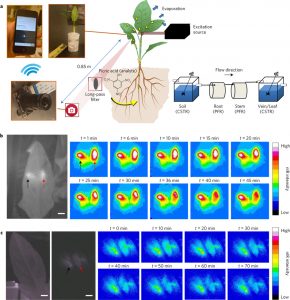 From these engineers’ perspective, plants are “microfluidic, self-powered autosamplers of their surroundings”. Wong et al. exploit this property to produce wild-type plants that sense and signal the presence of nitroaromatic contaminants (such as are found in explosives) in the soil. They infiltrate into leaf tissues single-walled carbon nanotubes conjugated to a peptide sensitive to the nitroaromatic picric acid. Picric acid binding leads to a conformation change that is translated to spectral modulations of the carbon nanotube’s fluorescence in the near infrared region, which can be recorded by a nearby photodetector. The authors observe that this nanobionic sensing approach can be extended to the detection of other compounds in “cities, crop fields, high-security facilities, and homes.” Nature Materials 10.1038/nmat4771 Tags: Applied Plant Biology, Biochemistry, Environmental Plant Biology
From these engineers’ perspective, plants are “microfluidic, self-powered autosamplers of their surroundings”. Wong et al. exploit this property to produce wild-type plants that sense and signal the presence of nitroaromatic contaminants (such as are found in explosives) in the soil. They infiltrate into leaf tissues single-walled carbon nanotubes conjugated to a peptide sensitive to the nitroaromatic picric acid. Picric acid binding leads to a conformation change that is translated to spectral modulations of the carbon nanotube’s fluorescence in the near infrared region, which can be recorded by a nearby photodetector. The authors observe that this nanobionic sensing approach can be extended to the detection of other compounds in “cities, crop fields, high-security facilities, and homes.” Nature Materials 10.1038/nmat4771 Tags: Applied Plant Biology, Biochemistry, Environmental Plant Biology
We like this paper so much that we’re including a video summary of it, enjoy! (And watch out, your plants may be spying on you….)
Have you spotted a new paper that you think will be of broad interest to the community? Feel free to share your finds below, or drop us an email!
