Karrikins are small molecules found in smoke that promote seed germination and have been 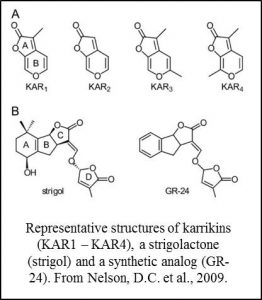 associated with the activation of seeds following fire (Nelson et al., 2012). Strigolactones are structurally similar small molecules that act as endogenous hormones and as secreted signals that promote the germination of parasitic plants and branching responses in mycrorrhizal fungi (Brewer et al., 2013; de Saint Germain et al, 2013; Waldie et al., 2014). Studies in Arabidopsis showed that these two families of molecules share some aspects of their recognition pathways. Specifically, the related proteins D14 and KAI2 are components of receptor complexes for strigolactones and karrikins respectively (Waters et al., 2012; Smith and Li, 2014).
associated with the activation of seeds following fire (Nelson et al., 2012). Strigolactones are structurally similar small molecules that act as endogenous hormones and as secreted signals that promote the germination of parasitic plants and branching responses in mycrorrhizal fungi (Brewer et al., 2013; de Saint Germain et al, 2013; Waldie et al., 2014). Studies in Arabidopsis showed that these two families of molecules share some aspects of their recognition pathways. Specifically, the related proteins D14 and KAI2 are components of receptor complexes for strigolactones and karrikins respectively (Waters et al., 2012; Smith and Li, 2014).
Phylogenetic studies indi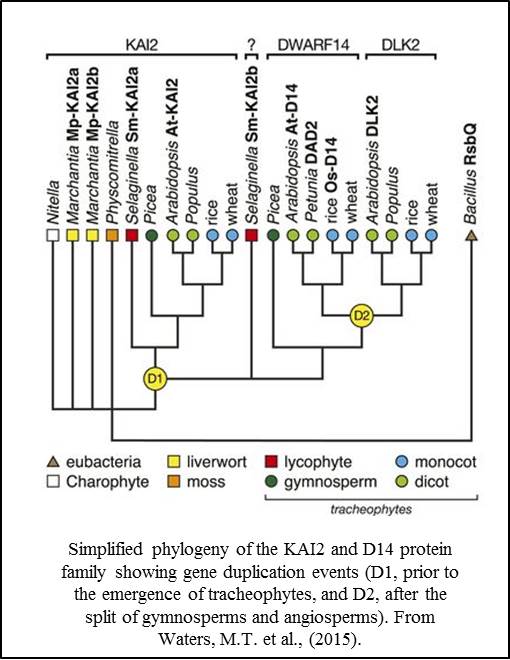 cate that the KAI2/D14 protein family has resulted from two or more gene duplication events. The KAI2 clade is thought to be ancestral, with a duplication event (D1) leading to the D14 clade and a second event (D2) occurring in angiosperms. Using a functional approach, Waters et al., (2015) showed that the KAI2-like genes in liverworts (Marchantia polymorpha) and lycophytes (Selaginella moellendorffii) could not substitute for D14 activity in Arabidopsis, but that one of the KAI2-like genes from Selaginella was able to partially substitute for KAI2 activity. These results support the model that KAI2 function is ancestral to D14 function, and that strigolactone perception by D14 proteins is derived.
cate that the KAI2/D14 protein family has resulted from two or more gene duplication events. The KAI2 clade is thought to be ancestral, with a duplication event (D1) leading to the D14 clade and a second event (D2) occurring in angiosperms. Using a functional approach, Waters et al., (2015) showed that the KAI2-like genes in liverworts (Marchantia polymorpha) and lycophytes (Selaginella moellendorffii) could not substitute for D14 activity in Arabidopsis, but that one of the KAI2-like genes from Selaginella was able to partially substitute for KAI2 activity. These results support the model that KAI2 function is ancestral to D14 function, and that strigolactone perception by D14 proteins is derived.
Additional studies have demonstrated another gene amplification event specifically occurring in parasitic plants of the Orobanchaceae family. Interestingly, this event, which is correlated with a germination response to strigolactones, occurred in the KAI2 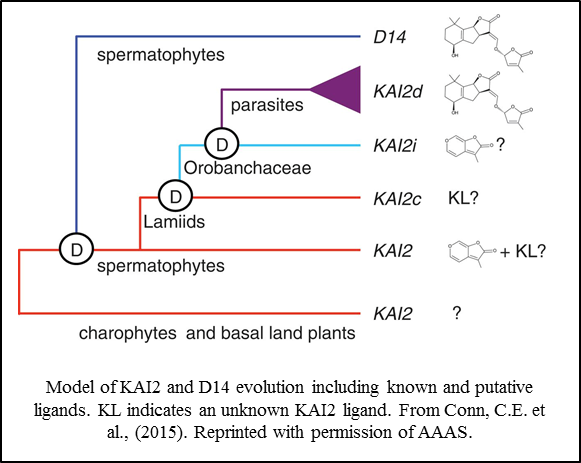 lineage, not the D14 lineage as might have been expected (Conn et al., 2015; Tsuchiya et al., 2015). These observations suggest that strigolactone perception arose twice independently within the KAI2/D14 protein family. The identity of preferred ligands for other members of the KAI2 protein family, including those expressed in green algae and basal land plants, remains obscure.
lineage, not the D14 lineage as might have been expected (Conn et al., 2015; Tsuchiya et al., 2015). These observations suggest that strigolactone perception arose twice independently within the KAI2/D14 protein family. The identity of preferred ligands for other members of the KAI2 protein family, including those expressed in green algae and basal land plants, remains obscure.
Open questions include the nature of the ancestral functions of the KAI2/D14 pathway in plants and green algae (see Delaux et al., 2012), the role of protein degradation (see Jiang et al., 2013; Zhou et al., 2013) and substrate hydrolysis in downstream signaling (see Tsuchiya et al., 2015), how endogenous strigolactones exert their diverse roles in plant development, and how fungi perceive strigolactones (see Ruyter-Spira, C. and Bouwmeester, H., 2012).
References
Brewer, P.B., Koltai, H., and Beveridge, C.A. (2013). Diverse roles of strigolactones in plant development. Mol Plant. 6: 18-28.
Conn, C.E., Bythell-Douglas, R., Neumann, D., Yoshida, S., Whittington, B., Westwood, J.H., Shirasu, K., Bond, C.S., Dyer, K.A. and Nelson, D.C. (2015). Convergent evolution of strigolactone perception enabled host detection in parasitic plants. Science. 349: 540-543.
de Saint Germain, A., Bonhomme, S., Boyer, F.D., and Rameau, C. (2013). Novel insights into strigolactone distribution and signalling. Curr Opin Plant Biol. 16: 583-589.
Delaux, P.M. et al., (2012). Origin of strigolactones in the green lineage. New Phyt. 195: 857–871.
Jiang, L., Liu, X., Xiong, G., Liu, H., Chen, F., Wang, L., Meng, X., Liu, G., Yu, H., Yuan, Y., Yi, W., Zhao, L., Ma, H., He, Y., Wu, Z., Melcher, K., Qian, Q., Xu, H.E., Wang, Y. and Li, J. (2013). DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature. 504: 401-405.
Nelson, D.C., Flematti, G.R., Ghisalberti, E.L., Dixon, K.W. and Smith, S.M. (2012). Regulation of seed germination and seedling growth by chemical signals from burning vegetation. Annu. Rev. Plant Biol. 63: 107-130.
Ruyter-Spira, C. and Bouwmeester, H. (2012). Strigolactones affect development in primitive plants. The missing link between plants and arbuscular mycorrhizal fungi?. New Phyt. 195: 730–733.
Smith, S.M., and Li, J. (2014). Signalling and responses to strigolactones and karrikins. Curr. Opin. Plant Biol. 21: 23 – 29.
Tsuchiya, Y., Yoshimura, M., Sato, Y., Kuwata, K., Toh, S., Holbrook-Smith, D., Zhang, H., McCourt, P., Itami, K., Kinoshita, T. and Hagihara, S. (2015). Probing strigolactone receptors in Striga hermonthica with fluorescence. Science. 349: 864-868.
Waldie, T., McCulloch, H., and Leyser, O. (2014). Strigolactones and the control of plant development: lessons from shoot branching. Plant J. 79: 607-622.
Waters, M.T., Nelson, D.C., Scaffidi, A., Flematti, G.R., Sun, Y.K., Dixon, K.W. and Smith, S.M. (2012). Specialisation within the DWARF14 protein family confers distinct responses to karrikins and strigolactones in Arabidopsis. Development. 139: 1285-1295.
Waters, M.T., Scaffidi, A., Moulin, S.L.Y., Sun, Y.K., Flematti, G.R. and Smith, S.M. (2015). A Selaginella moellendorffii ortholog of KARRIKIN INSENSITIVE2 functions in Arabidopsis development but cannot mediate responses to karrikins or strigolactones. Plant Cell. 27: 1925-1944.
Zhou, F., Lin, Q., Zhu, L., Ren, Y., Zhou, K., Shabek, N., Wu, F., Mao, H., Dong, W., Gan, L., Ma, W., Gao, H., Chen, J., Yang, C., Wang, D., Tan, J., Zhang, X., Guo, X., Wang, J., Jiang, L., Liu, X., Chen, W., Chu, J., Yan, C., Ueno, K., Ito, S., Asami, T., Cheng, Z., Wang, J., Lei, C., Zhai, H., Wu, C., Wang, H., Zheng, N., and Wan, J. (2013). D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 504: 406-410.
This post is part of our “Research in Focus” series, highlighting significant plant science breakthroughs and discoveries from around the globe, by Mary Williams (@PlantTeaching). If you have ideas or suggestions please email mwilliams@aspb.org, using Research in Focus as the subject, and include a link as well as an explanation of why this should be highlighted.
